I’m trying to figure out why I get very consistently different results when doing NMF with Librosa vs with FluCoMa. I bet someone here knows the answer. @weefuzzy ? @tremblap ? @jamesbradbury
I’m trying to be as similar as possible in the execution but continuously get these different results (code below). This is the first three seconds of the FluCoMa test soundfile “Tremblay-BaB-SoundscapeGolcarWithDog.wav”.
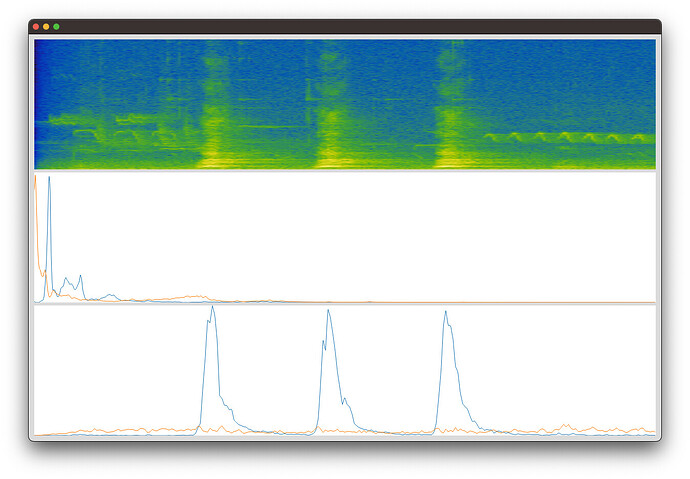
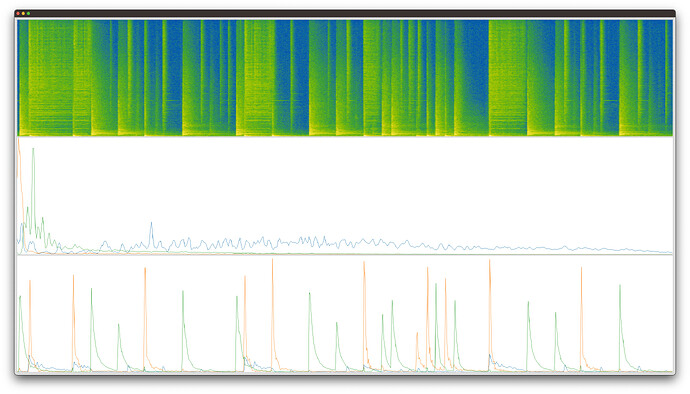
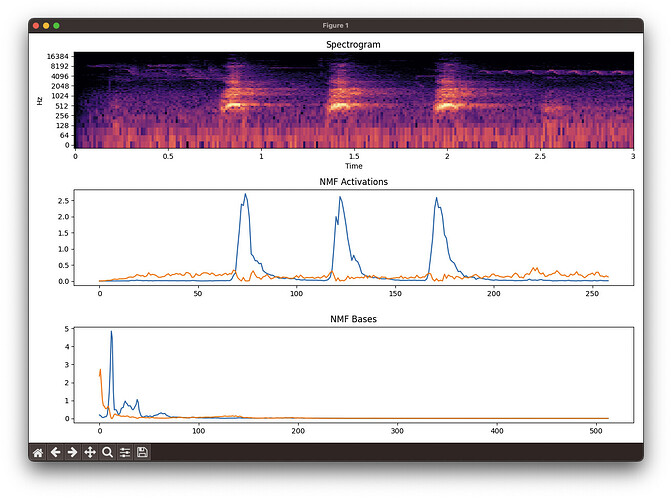
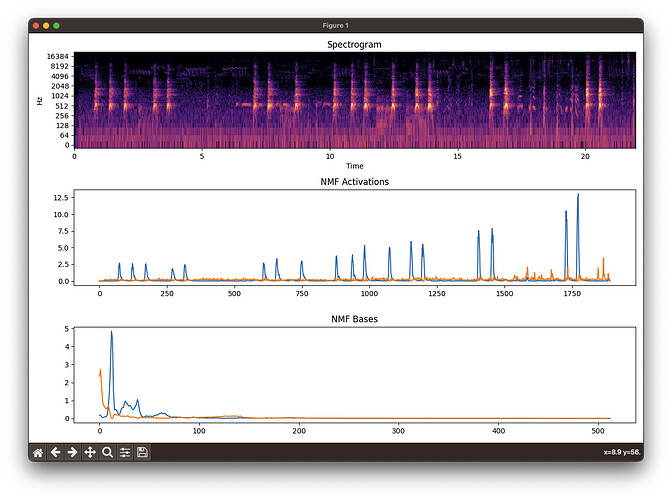
FluCoMa (spectrogram, bases, activations):
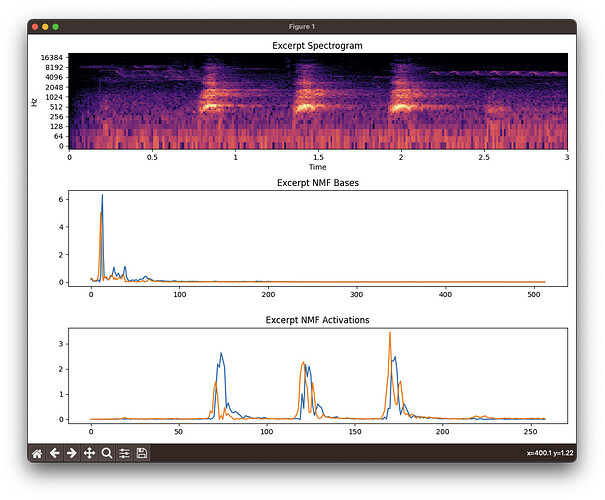
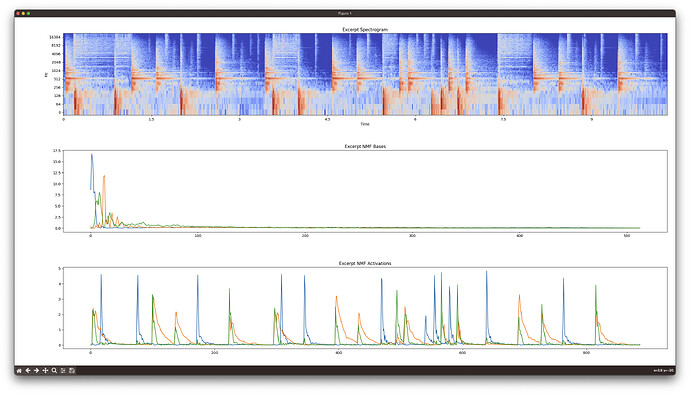
Librosa (spectrogram (log by default), bases, activations):
You can see that FluCoMa consistently separates out the dog bark (blue basis and activation) from the “other” in the sound file. Librosa does not.
Audio Results:
I’ve also resynthesized the buffer in Librosa and FluCoMa to aurally compare:
https://drive.google.com/drive/folders/1cx8wDh9DNhH2GoIpsbpaJzPtOg5PY7X0?usp=sharing
It’s obvious to me that FluCoMa works better (sounds better, separates better). Why is this? How could I get as good of results in Librosa or Python generally?
Code:
FluCoMa:
(
~thisFolder = thisProcess.nowExecutingPath.dirname;
fork{
b = Buffer.read(s,FluidFilesPath("Tremblay-BaB-SoundscapeGolcarWithDog.wav"),numFrames:44100*3);
~bases = Buffer(s);
~acts = Buffer(s);
~src = Buffer(s);
~resynth = Buffer(s);
s.sync;
2.do{
arg i;
FluidBufCompose.processBlocking(s,b,startChan:i,numChans:1,gain:0.5,destGain:1,destination:~src);
};
FluidBufNMF.processBlocking(s,~src,bases:~bases,resynth:~resynth,resynthMode:1,activations:~acts,components:2);
~mags = Buffer(s);
FluidBufSTFT.processBlocking(s,~src,magnitude:~mags,action:{"done".postln;});
~tmpBuf = Buffer(s);
2.do{
arg i;
FluidBufCompose.processBlocking(s,~resynth,startChan:i,numChans:1,destination:~tmpBuf);
s.sync;
~tmpBuf.write(~thisFolder+/+"flucoma-component-%.wav".format(i),"wav");
s.sync;
};
}
)
(
~win = Window();
~win.layout = VLayout(
FluidWaveform(imageBuffer:~mags,parent:~win,standalone:false,imageColorScheme:1,imageColorScaling:1),
FluidWaveform(featuresBuffer:~bases,parent:~win,standalone:false,normalizeFeaturesIndependently:false),
FluidWaveform(featuresBuffer:~acts,parent:~win,standalone:false,normalizeFeaturesIndependently:false)
);
~win.front;
)
Librosa:
import librosa
import librosa.display
import numpy as np
import matplotlib.pyplot as plt
import soundfile as sf
def resynthComponent(stft,basis,activation,outpath,fftSettings):
Y = np.outer(basis, activation) * np.exp(1j * np.angle(stft))
y = librosa.istft(Y,**fftSettings)
sf.write(outpath,y,sr,subtype='PCM_24')
# Replace 'your_audio_file.wav' with the path to your audio file
audio_file_path = '../audio-files/Tremblay-BaB-SoundscapeGolcarWithDog.wav'
n_components = 2
sr = 44100
fftSettings = {'n_fft':1024,'hop_length':512}
excerpt, sr = librosa.load(audio_file_path,sr=sr,duration=3,mono=True)
y, sr = librosa.load(audio_file_path,sr=sr)
print(audio_file_path,sr)
excerpt_stft = librosa.stft(excerpt,**fftSettings)
print(f'stft shape: {excerpt_stft.shape[0]} mags, {excerpt_stft.shape[1]} frames')
bases, activations = librosa.decompose.decompose(np.abs(excerpt_stft), n_components=n_components)
bases = np.transpose(bases)
print(f'bases shape: {bases.shape}')
print(f'activations shape: {activations.shape}')
for i in range(n_components):
resynthComponent(excerpt_stft,bases[i],activations[i],f'librosa-component-{i}.wav',fftSettings)
plt.figure(figsize=(12, 8))
plt.subplot(3, 1, 1)
librosa.display.specshow(librosa.amplitude_to_db(np.abs(excerpt_stft)), sr=sr, x_axis='time', y_axis='log')
plt.title('Excerpt Spectrogram')
plt.subplot(3, 1, 2)
for basis in bases:
plt.plot(basis)
plt.title('Excerpt NMF Bases')
plt.subplot(3, 1, 3)
for activation in activations:
plt.plot(activation)
plt.title('Excerpt NMF Activations')
plt.tight_layout()
plt.show()